We have compiled tips to void the 6 most common pitfalls of cytometry experiment design and analysis. Learn how to avoid failed experiments, save time and frustration and get better data!
Sections:
1. Reagent/Fluorochrome Selection
According to our survey of core managers, the most common mistake when designing a multicolor flow cytometry panel is failing to consider how each fluorescent antibody will work on your particular cytometer. FluoroFinder takes the guesswork out of panel building by preloading your cytometer configurations and dynamically eliminating reagent/fluorochrome combinations that won’t work for you. Furthermore, you can compare spectral properties for reagents from any supplier on FluoroFinder’s unique spectra viewer to ensure that they will work together on your machine. Find your core lab, here.
2. Controls
Another commonly reported mistake in fluorescent experiment design is the lack of proper controls. Be sure you include all the necessary controls for your experiment type, including:
-
- Viability Dyes – Viability dyes allow you to distinguish between live and dead cells in your sample population. This is important as dead cells can significantly skew your results One of the biggest challenges is determining which dyes will work with your specific lasers and optical filters. With FluoroFinder, you can automatically view all the available and optimal viability dyes for your cytometer and view each dye’s spectral profile to determine how it will fit with the rest of your panel.
- FMO – Used to determine proper gating of a multicolor panel, FMO controls contain all the fluorochromes in a panel, minus the one being measured. This allows you to assess the spread of all the fluorophores into the missing channel and then set up gates accordingly.
- Isotype – Isotype-specific controls are useful to determine if an antibody in your panel shows non-specific binding affinity for your cells.
- Dump Channel – Also called an exclusion channel, a dump channel is used to gate out any cells that are not of interest. For example, a dump channel can be combined with a viability dye to exclude dead cells from analysis.
3. Sample Preparation
Experienced flow core managers report seeing poorly prepared samples as the most common mistake for new flow researchers. Before running a sample through your cytometer, cells must be properly suspended, filtered and de-aggregated for successful analysis. For a detailed list of recommended sample prep techniques, please see our pre-sort checklist article.
4. Spillover
Another very common mistake in panel design is choosing reagents with overlapping emission spectra that can cause false signals and affect your analysis. Therefore, it is important to design your panels with reagents that minimize this spectral “spillover”. Researchers can use FluoroFinder’s SpectraViewer to compare reagents from multiple vendors to see how they will work together and have confidence in their reagent selections. Sample of a FluoroFinder spectra viewer showing potential spillover issue with the yellow/red fluors.
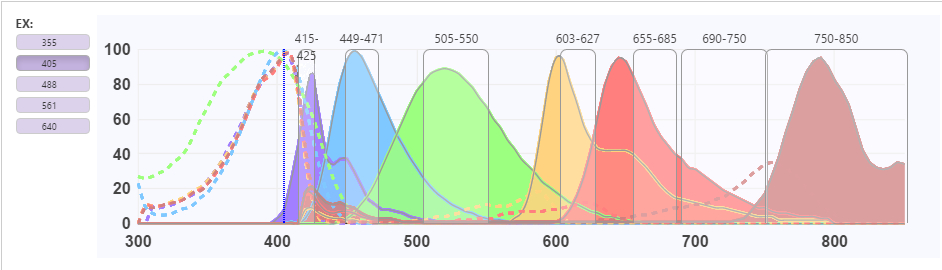
5. Compensation
Even with the best-designed panel, you are still likely to experience some spillover. Compensation is the process for correcting for spillover, however, it is also a major source of experimental mistakes in flow cytometry. While it may be tempting to compensate your data manually based on how the data looks, this practice often leads to overcompensation and faulty data. Therefore, it is strongly recommended that you use an automatic compensation algorithm supplied with your cytometer analysis software. If you are unsure about compensating with your analysis software, consult your core manager for their recommendations.
6. Gating
The final most common mistake in cytometry analysis is improper gating. Gating is the process of setting boundaries to define subsets of events. Gates can be used for either data acquisition or analysis by drawing boundaries around subsets on histograms or dot data plots. Like compensation, gates should never be set manually based on how the data looks. Gates should be set based on the boundaries of positivity determined by FMO controls to ensure that only true positive cells are sorted.





